
singlet¶
Single cell RNA-Seq analysis with quantitative phenotypes.
Requirements¶
Python 3.4+ is required. Moreover, you will need:
- pyyaml
- numpy
- scipy
- pandas
- xarray
- scikit-learn
- matplotlib
- seaborn
Install¶
To get the latest stable version, use pip:
pip install singlet
To get the latest development version, clone the git repo and then call:
python3 setup.py install
Usage example¶
You can have a look inside the test folder for examples. To start using the example dataset:
- Set the environment variable SINGLET_CONFIG_FILENAME to the location of the example YAML file
- Open a Python/IPython shell and type:
from singlet.dataset import Dataset
ds = Dataset(
samplesheet='example_sheet_tsv',
counts_table='example_table_tsv')
ds.counts = ds.counts.iloc[:200]
vs = ds.dimensionality.tsne(
n_dims=2,
transform='log10',
theta=0.5,
perplexity=0.8)
ax = ds.plot.scatter_reduced_samples(
vs,
color_by='quantitative_phenotype_1_[A.U.]')
plt.show()
This will calculate a t-SNE embedding of the first 200 features and then show your samples in the reduced space. It should look like this:
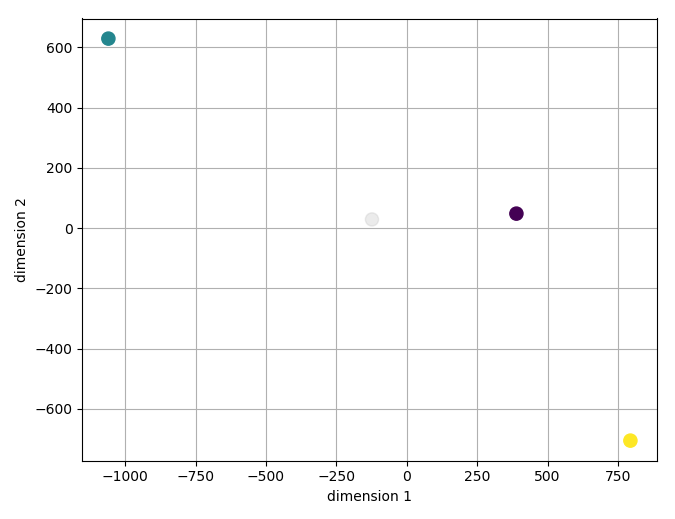
Note
The figure looks different on OSX, but no worries, if you got there without errors chances are all is working correctly!
